The Post That Broke the Science Feed
A post starts circulating on X. A software engineer in Sydney built a cancer vaccine for his dying dog using AI tools and AlphaFold. The tumor shrank. Biomedical researchers, AI optimists, and journalists all pick it up.
So what actually happened?

What’s True, What’s Overstated
- ✓ Paul Conyngham is a software engineer, not a biomedical researcher. He’s a Sydney-based software developer with no formal training in oncology, biochemistry, or vaccine design.
- ~ He used AI to design the vaccine. True, and then some. Multiple AI systems were involved across different stages of the pipeline — not a single tool, not a single prompt. The AI did the heavy lifting on neoantigen candidate selection and construct design; Conyngham directed it.
- ~ He used AlphaFold to design the vaccine. Partially true. AlphaFold was used to model protein structures — specifically to predict the 3D conformation of candidate neoantigens and assess whether mutations meaningfully altered surface exposure. AlphaFold does not identify mutations, call variants from sequencing, or design vaccines. It contributed structural validation after candidate neoantigens were identified through other means.
- ✗ He synthesized it himself. mRNA synthesis requires laboratory equipment that isn’t available in a home setting. Professor Pall Thordarson and his team at the University of New South Wales synthesized the mRNA construct. Conyngham did the informatics pipeline; UNSW did the wet lab.
- ~ The tumor shrank by 50%. Wrong number. Reports that followed the story more carefully put the tumor reduction at 75%, not 50%. The actual result is better than the viral version.
- ✓ The dog had a non-responding sarcoma. Rosie had a sarcoma that had not responded to conventional treatment. The vaccine wasn’t a first-line attempt — it was a last-resort intervention for a tumor that conventional oncology had already failed to control.
- ✓ This is AI enabling something that wasn’t previously accessible. The pipeline Conyngham assembled would have required a research team and years of work a decade ago. He did it in weeks as a solo engineer with AI assistance. That’s real and worth taking seriously.
The accurate version is more impressive than the viral one — a 75% tumor reduction in a non-responding case, built by one engineer with no biology background. The constraints (institutional manufacturing, professor collaboration, months of work) don’t diminish the result. They show you exactly where the frontier is.
What Actually Happened
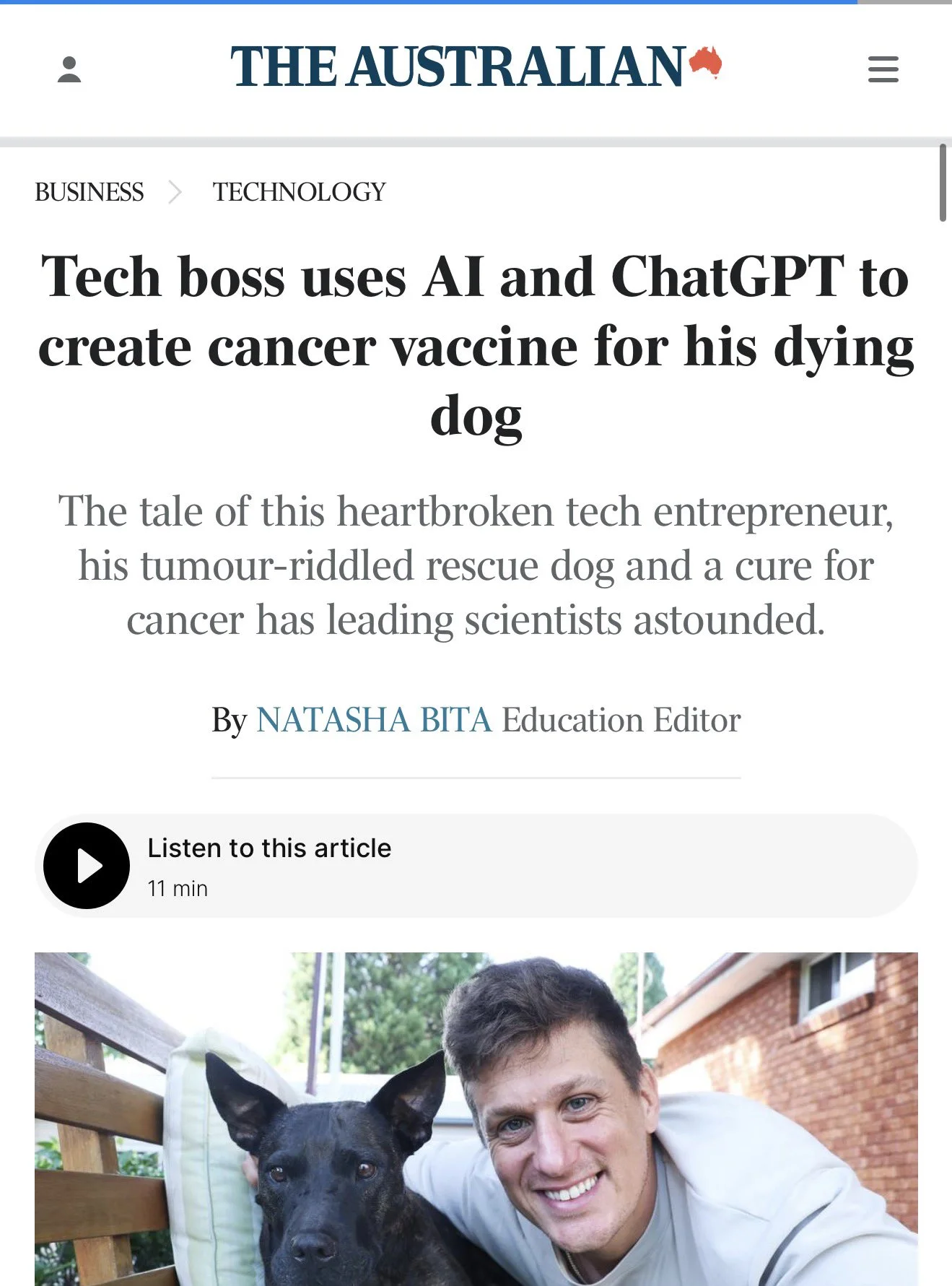
Rosie was diagnosed with a soft tissue sarcoma. After conventional treatments stalled — chemotherapy and surgery hadn’t stopped the tumor’s progression — Conyngham started researching personalized cancer vaccines.
This isn’t fringe science. Personalized neoantigen vaccines are one of the most active areas in oncology right now. The basic idea: every tumor accumulates mutations that healthy cells don’t have. Some of those mutations produce altered peptides — neoantigens — that the immune system can recognize as foreign. A vaccine that trains the immune system to target those specific peptides kills tumor cells while leaving normal tissue alone. The hard part is figuring out which mutations produce good neoantigens, and doing it fast enough to matter.
Conyngham ran Rosie’s tumor and blood through whole-exome sequencing to identify somatic mutations — changes in the tumor that normal cells don’t have. He used AI tools to run neoantigen prediction: filtering candidates by binding affinity to Rosie’s MHC alleles (the immune system’s “display system”), stability, and immunogenic potential. AlphaFold came in at the structural validation step — modeling the 3D shape of candidate neoantigens to check whether the mutations created surface-exposed changes a T-cell could recognize.
The resulting mRNA construct design was taken to Professor Pall Thordarson at UNSW, where the vaccine was synthesized and formulated. Rosie received it. The tumor reduced by approximately 75%.


This is not a peer-reviewed clinical result. It’s an n=1 case with no controls. But it’s also not nothing. The tumor that wasn’t responding to anything started responding to this. That’s data.
The Science: How Personalized Neoantigen mRNA Vaccines Work
Understanding why this works — and why it’s still hard — takes a short detour into cancer immunology.
Tumors Are Genetically Distinct From You
Every cancer is, functionally, a foreign organism. As cells divide and mutate, they accumulate genetic changes that healthy cells don’t have. Some of those changes alter the proteins those cells produce. When altered proteins are degraded inside the cell, fragments (peptides) get loaded onto MHC molecules and displayed on the cell surface — the immune system’s way of checking that a cell is “self.”
A mutation that changes a peptide sequence can create something the immune system has never seen before: a neoantigen. If the immune system can learn to recognize it, it can target and kill cells displaying it — which means tumor cells, not normal cells.
This is the conceptual foundation for personalized cancer vaccines. The challenge is identifying which of a tumor’s thousands of mutations produce neoantigens that:
- Actually get displayed on MHC (not all peptides do)
- Bind well enough to MHC to be stable (binding affinity)
- Can be recognized by T-cells (immunogenicity)
- Are absent from normal tissue (tumor-specificity)
The computational tools for this have gotten very good in the last five years. Not perfect. Good enough to produce candidate lists worth acting on.
Why mRNA
The vaccine format — mRNA — matters. mRNA vaccines encode the neoantigens directly. You inject a lipid nanoparticle carrying the mRNA sequence; cells take it up, produce the neoantigen peptides, present them on MHC, and the immune system develops a targeted response. No live virus, no viral vector, no permanent DNA modification.
The manufacturing complexity is real — mRNA synthesis, purification, and LNP formulation require controlled lab conditions. But the design step (which peptides to encode) is now largely computational, which is what makes the Conyngham story possible.
Where AlphaFold Actually Fits
AlphaFold predicts protein structure. Given a peptide sequence, it can model the 3D shape with high accuracy. In neoantigen pipeline terms, this is useful for:
- Checking whether a mutation creates a structurally distinct surface that could be immunogenic
- Assessing peptide-MHC binding geometry for borderline candidates
- Ruling out candidates where the mutation is buried inside the protein and won’t be displayed
AlphaFold did not “design” Rosie’s vaccine. It contributed structural evidence for candidate selection. That’s a supporting role, not the lead — but it’s a legitimate one.
Why This Matters Beyond One Dog
Rosie’s story isn’t happening in isolation. The personalized neoantigen vaccine space has been building serious clinical evidence.
Moderna and Merck’s mRNA-4157 (now called V940) is a personalized neoantigen vaccine for melanoma. In Phase 2b results published in 2023, combining it with Keytruda reduced the risk of recurrence or death by 49% compared to Keytruda alone. Phase 3 trials are now running. This is real pharmaceutical development, not a startup pitch.
In 2023, a Memorial Sloan Kettering team published results in Nature for personalized neoantigen vaccines in pancreatic cancer — one of the hardest-to-treat malignancies — showing that patients who mounted an immune response remained disease-free significantly longer than those who didn’t. Small trial, but the mechanistic signal was clear.
And for dogs specifically: Merck received USDA conditional approval in 2024 for a personalized mRNA cancer vaccine for canine osteosarcoma (bone cancer). The regulatory pathway for companion animal cancer is notably more accessible than human clinical approval, which is part of why canine oncology is becoming a proving ground for personalized vaccine approaches.
Conyngham’s work sits in this context. It’s not a lone genius moment — it’s one engineer finding his way to a methodology that serious research institutions are validating in parallel.
How to Save Lassie: A Non-Engineer’s Guide to the Same Pipeline

This is the part the viral post skipped.
The claim buried in the coverage — “AI has democratized cancer vaccines” — is half right. The informatics pipeline is now accessible to someone with patience and AI assistance, no formal background required. The manufacturing step is not. Knowing where the line falls is the difference between being inspired and being deluded.
Here’s what the pipeline looks like, step by step, from the perspective of someone with no bioinformatics training.
Step 1: Get the Sequencing Done
You need two samples from your dog (or yourself, or whoever you’re trying to help):
- A tumor sample — typically from a biopsy or surgical resection
- A normal sample — blood is standard
Both go through whole-exome sequencing (WES) or whole-genome sequencing (WGS). WES targets the protein-coding regions and is cheaper; WGS covers the full genome and gives more data. For neoantigen work, WES is usually sufficient.
Cost: roughly $300–$1,000 for WES through a commercial lab, or $500–$5,000 for WGS depending on depth. Companies like Illumina partners, Genoptix, and academic sequencing cores will do this. You need a veterinarian or physician to collect and submit the samples properly.
What you get back: raw sequencing data (FASTQ files), enormous by default. This is where the complexity starts, but also where AI assistance starts earning its keep.
Step 2: Variant Calling — Finding the Mutations
Variant calling compares your tumor sequencing to your normal sequencing to identify somatic mutations — changes in the tumor that aren’t in normal cells.
The standard open-source tool for this is GATK (Genome Analysis Toolkit) from the Broad Institute. Running it requires bioinformatics infrastructure — cloud compute, some Linux command-line work — but it’s well-documented, and AI assistants can walk you through every step.
This step will take time. Days, probably. Learning the tools, running pipelines, debugging file format issues. The tools assume you know bioinformatics conventions (BAM files, VCF format, reference genomes). An LLM can explain every concept, generate every command, and debug every error — but you still have to do the learning.
Output: a VCF file listing somatic mutations. Hundreds to thousands of entries.
Step 3: HLA Typing — Your Dog’s Immune System’s Vocabulary
MHC molecules (called HLA in humans, DLA in dogs) are the display system the immune system uses to show peptides to T-cells. Different individuals have different HLA alleles, and peptides bind differently to different alleles. You need to know which HLA alleles your patient has before you can predict which neoantigens will be displayed.
HLA typing is done from the normal sequencing data you already have. Tools: OptiType (free, open-source) for Class I, HLA-HD for Class I and II. Both run on standard Linux systems.
AI assistance here: feed the tools’ documentation to your LLM, explain your situation, and follow the guided setup. The commands are not intuitive, but they’re learnable.
Step 4: Neoantigen Prediction
This is the core of the pipeline. Given your mutation list and HLA alleles, you want to predict which mutations produce peptides that:
- Bind to the patient’s HLA alleles (predicted by binding affinity models)
- Are likely to be immunogenic
- Are absent from normal tissue
The two main tools:
pVACtools (Washington University, St. Louis) — an open-source neoantigen prediction pipeline that integrates multiple binding prediction algorithms. It takes your VCF file and HLA types, runs the predictions, and outputs a ranked candidate list with binding affinity scores, expression data (if you have RNA-seq), and tumor variant allele frequency. The documentation is good. The GitHub repo is actively maintained. This is what serious neoantigen research groups use.
NetMHCpan — a binding affinity prediction tool from DTU (Technical University of Denmark). Available as a web server for small queries, downloadable for larger runs. pVACtools calls it internally; you don’t necessarily need to run it separately.
What you get: a ranked list of candidate neoantigens, typically filtered to those with binding affinity below a threshold (strong binders: IC50 < 500nM, weak binders: < 1000nM).
Step 5: Structural Validation with AlphaFold
This is where Conyngham used AlphaFold. For your top candidates — the peptides with the best predicted binding affinities — you can model the 3D structure of the mutant peptide versus the wildtype to see whether the mutation creates a surface-exposed change.
AlphaFold 3 is available as a web server (alphafoldserver.com). You can submit peptide sequences and get structure predictions in minutes, for free.
In practice: for each candidate neoantigen, compare the predicted structure of the mutant peptide-MHC complex to the wildtype. If the mutation is surface-exposed and creates a distinct structural feature, it’s a stronger candidate. If it’s buried inside the protein, deprioritize it.
An LLM can help you interpret the structural outputs and think through the selection criteria. Not for the computation — for helping a non-expert reason about what the computation is telling them.
Step 6: mRNA Construct Design
Once you have your finalized candidate list (typically 10–20 neoantigens for a personalized vaccine), you need to design the mRNA construct — the sequence encoding those peptides, optimized for expression.
This involves:
- Codon optimization: rewriting the coding sequence to use codons (DNA triplets) that produce efficient translation in mammalian cells. Tools: GENEWIZ’s codon optimizer, Geneious, or simply prompting an LLM with the relevant constraints.
- UTR selection: 5’ and 3’ untranslated regions that affect mRNA stability and translation efficiency. Standard UTR sequences from published literature are available; ask an LLM to point you to the current best practices.
- Poly-A tail: included at the 3’ end for stability. Standard protocol.
This is where AI does the most work — reasoning about construct design choices, interpreting literature, pulling recommendations from multiple sources into a coherent sequence.
Output: a DNA sequence encoding your construct, ready for synthesis.
Step 7: The Hard Part — Manufacturing
This is where you need institutional help. Full stop.
mRNA synthesis requires: - In vitro transcription (IVT) with purified enzymes and template DNA - mRNA capping (for stability and translation) - Purification (HPLC or similar) - Formulation into lipid nanoparticles (LNP) for delivery - Sterility and endotoxin testing before administration
None of this is doable at home. The equipment costs hundreds of thousands of dollars. The reagents require institutional procurement. The safety testing is non-negotiable.

What Conyngham did — find a university professor willing to help with synthesis — is the realistic path. Professors with relevant lab infrastructure exist. Academic medical centers, veterinary schools, and research hospitals often have capacity and interest in interesting applications. You need to find someone, explain what you’ve built, and ask.
This is not easy. But it’s not impossible either. The informatics work Conyngham did — producing a credible candidate list with a properly documented pipeline — is what makes the ask to a professor go from “can you help my sick dog” to “here’s a pipeline I’ve run, here are the top 15 candidate neoantigens with supporting structural data, can you synthesize this construct.”
That’s a different conversation.
Realistic Timeline and What Hard Looks Like
From sequencing results to a synthesized vaccine: 3–6 months if you’re working on this seriously alongside other responsibilities, with AI assistance, and assuming you find a willing lab partner. The pipeline steps aren’t individually difficult — each one is learnable — but there are a lot of them, the failure modes aren’t always obvious, and the learning curve in bioinformatics is real.
What will block you: - VCF file format errors in variant calling (GATK is particular) - HLA typing tools failing on non-standard canine reference genomes (dog HLA databases are less complete than human) - pVACtools dependency installation (Python environment management, library conflicts) - Finding a lab partner willing to synthesize and administer
What won’t block you: - Not having a biology degree - Not understanding the math behind binding affinity models - Not knowing what a FASTQ file is at the start
The LLM fills those gaps. Not perfectly, not always correctly — you have to verify its output, exactly as Conyngham did. But the ability to ask “what does this pVACtools output mean” and get a coherent answer is a real capability that didn’t exist five years ago.
What This Actually Signals

The Rosie story matters for a reason that’s easy to miss in the breathless coverage: it’s a proof of concept for pipeline accessibility, not a proof of concept for the underlying science.
The underlying science — personalized neoantigen mRNA vaccines — is already validated. Moderna’s Phase 3 trial for melanoma is not waiting on a Sydney engineer to prove the concept. That evidence is building independently, in large clinical trials, with rigorous endpoints.
What Conyngham showed is that the informatics portion of this pipeline has crossed an accessibility threshold. A motivated non-expert, with AI assistance, can now run neoantigen prediction with the same tools academic researchers use. That didn’t take an AI breakthrough. It took good open-source tools (pVACtools, AlphaFold), LLMs that can explain complex software, and sequencing that costs less than a used laptop.
The manufacturing barrier is real and isn’t going away. You need institutional help. But the information barrier — which used to be the bigger bottleneck — has dropped hard.
That’s what’s worth paying attention to. Not “AI made a cancer vaccine” — but “AI made the hard part of the pipeline accessible to people who couldn’t get there before.” The hard part used to be knowing which mutations mattered. Now it’s getting access to a lab.
That’s a different problem. And it’s a more tractable one.